Antibiotic resistance of bacteria and other pathogens which renders various drugs useless has become a threat that is progressing at an alarming rate. This hazard has reached a point where scientists estimate that these superbugs could kill 10 million people a year if we can’t find a solution. The discovery of antibiotics earlier was a blessing for mankind. However, over the years, due to incorrect prescription, patients not completing required doses, and especially self-medication of drugs available over-the-counter, among other reasons, has resulted in antibiotic resistance.
The problem is that doctors cannot immediately tell by diagnosis whether the patient visiting their clinic has an infection from microbes which are resistant to antibiotics, or which antibiotics. To confirm this, they have to send samples to a testing lab which would give the results after one to two days.
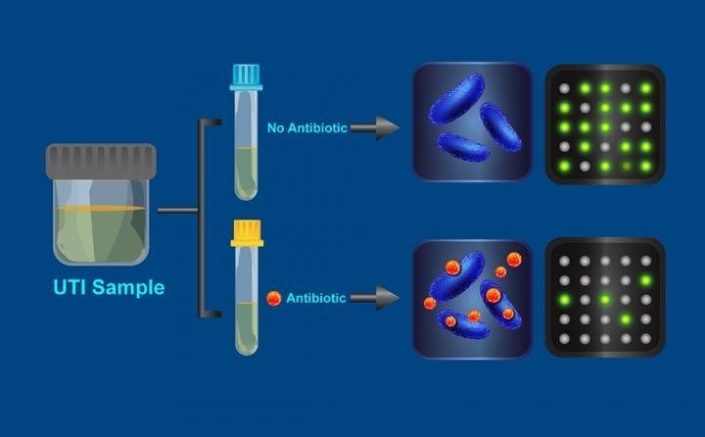
In the new test, a sample of urine containing bacteria from a UTI is divided in half, with one part being exposed to an antibiotic: then, if the bacteria is immune to that antibiotic, it should have the same number of fluorescent spots as the untreated sample
(Source: http://www.caltech.edu/news/caltech-researchers-create-test-reveals-antibiotic-resistant-bacteria-30-minutes-79943)
A new test developed at Caltech University identifies antibiotic-resistant bacteria in 30 minutes, whose study was published in the journal of Science Translational Medicine. To develop the test, the team focused on urinary tract infections (UTIs), one of the most common types of bacterial infection. To detect the resistance of any present bacteria, a doctor first collects a urine sample from a patient with a UTI. They treat half of the sample with an antibiotic for 15 minutes, while the other half is tested without adding any drugs. After this, a process that combines a chemistry detection technique called dLAMP, with a device called SlipChip, searches for specific DNA markers.
The SlipChip device displays the biomarkers as fluorescent spots. A less number of spots indicate that the antibiotic was more effective against the bacteria in the sample. If the antibiotic-treated and non-treated samples have similar numbers of fluorescent spots, the drug clearly had no effect. But if considerably fewer spots show up in the treated sample, then that antibiotic can be prescribed to the patient, since the bacteria is not resistant against it.
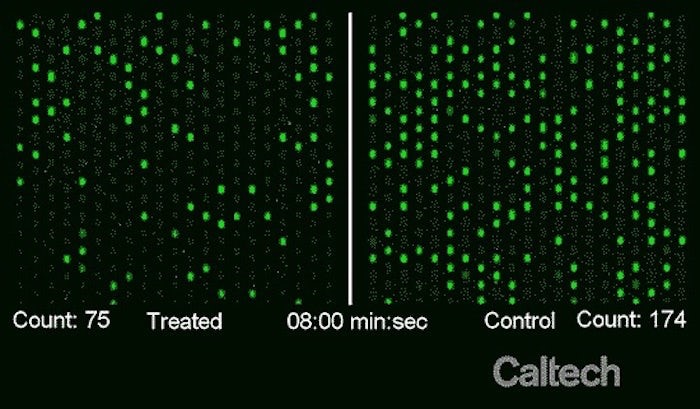
The treated sample (left) has far fewer fluorescent spots than the control sample on the right, indicating the antibiotic used was effective
(Source: http://www.caltech.edu/news/caltech-researchers-create-test-reveals-antibiotic-resistant-bacteria-30-minutes-79943)
The researchers tested this method on 54 urine samples from patients with Escherichia coli caused UTIs. They compared the test results with the standard test for detecting antibiotic resistance, which takes two days, and found that their results had an accuracy of around 95%.
When doctors treat bacterial infections, they usually skip first-line antibiotics like methicillin or amoxicillin, which are drugs that bacteria are more likely to be resistant to, and instead go ahead with stronger second-line antibiotics, like ciprofloxacin. This practice increases the efficiency of the treatment, but it is not ideal. The increased use of second-line antibiotics increases the chances of development of antibiotic resistance.
With this new testing method, hopefully, doctors will be able to prescribe antibiotics appropriately as per the resistance of the pathogen. Next, the team plans to further develop the test to detect other bacteria besides E. coli and develop the method to work with blood samples as well.
References:
1] Schoepp, N. G., Schlappi, & Ismagilov, R. F. (2017). Rapid pathogen-specific phenotypic antibiotic susceptibility testing using digital LAMP quantification in clinical samples. Science Translational Medicine, 9(410). doi:10.1126/scitranslmed.aal3693
2] http://www.pasadenanow.com/main/caltech-researchers-create-test-that-reveals-antibiotic-resistant-bacteria-in-30-minutes/#.WdYwt9hLdPa
3] http://newatlas.com/half-hour-test-superbugs/51628/






Thanks for sharing the information on antibiotic resistance bacteria detected in only 30 min.