RNA extraction using TRIzol
Materials:
Reagents
- TRIzol Reagent
- Chloroform
Set up requirements
Fumehood, vortexer, micropipette, chilled micro-centrifuge, pellet pestle homogenizer, and all standard equipment of molecular biology laboratory.
Precautions:
- Always use disposable nitrile gloves
- Use sterilize tips or RNAase/DNAase free filter tips
- Use disposable sterile plastic ware or RNAase free glassware
- Before starting the extraction, clean the workstation with surface decontamination that destroys RNAases (RNA-zap).
- Chill the micro-centrifuge to 4∞C
Procedure:
1. Homogenization
- Add 1ml of TRIzol per 100 mg of fresh tissue or tissue stored in RNA later and mince on ice using sterile scalpels. The tissue is further homogenized with a sterile pellet pestle probe. Frozen tissue should be thawed at room temp and removed from RNA later before adding TRIzol reagent.
2. Extraction
- Transfer the tissue lysate to a 2 ml sterile micro-centrifuge tube
- For complete dissociation of nucleoprotein complexes, let the homogenate settle at room temp for 5 min.
- Add 200μl of chilled Chloroform and vortex vigorously for 20 sec. Thorough mixing and shaking is critical for phase separation.
- Let the homogenate to settle at room temp for 5 min.
- Centrifuge the homogenate at 12,000 x g for 20 min at 4∞C.
3. Precipitation
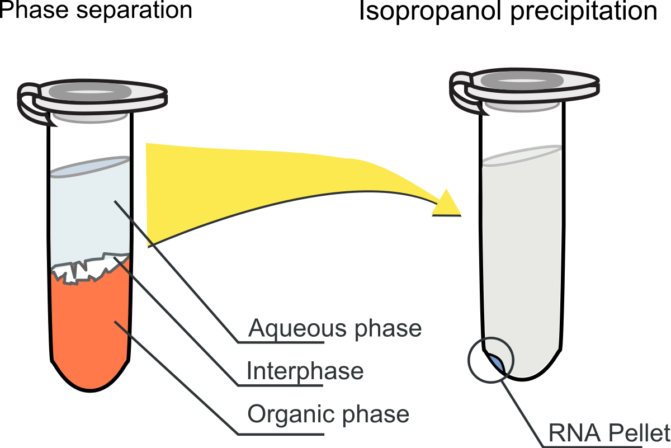
- Carefully transfer the upper aqueous phase (600μl) over 500 μl of chilled isopropanol in a microcentrifuge tube. Store interphase for further isolation of DNA.
- Let the RNA, to precipitate in isopropanol for 1 to 2 hr at -20∞C.
- After precipitation, centrifuge the RNA sample in isopropanol 15,000 x g for 20 min at 4∞C.
- Carefully extract the RNA precipitated in the form of a gel at the bottom of the microcentrifuge tube.
- RNA pellet so obtained is washed with 200μl of 75% ethanol proceeded with brief vortexing and
centrifugation at 7500 g for 10 min at 4°C. - Air-dried the washed RNA pellet for 5 min
- Using RNAase free water re-suspend the air-dried RNA pellet.
Now, you can proceed with RNA quantity and quality check, followed by its conversion into cDNA.


